Alla Krasikova

Personal information
Education & Qualifications
| May 2007 |
| PhD degree in Cytology, histology and cell biology (St. Petersburg State University, Faculty of Biology and Soil sciences, Department of Cytology and Histology). |
| |
September 1997-June 2003
|
| Master Sc. degree in Biophysics (Saint-Petersburg State Polytechnical University, The Faculty of Physics and Mechanics, Department of Biophysics, graduated with honors). |
Current Position
Institute: Saint-Petersburg State University, Department of Cytology and Histology, Laboratory of the cell nucleus structure and dynamics
Positions: (1) Head of laboratory, (2) Associate Professor (full-time).
The field of scientific interests: functional compartments of the cell nucleus, genome architecture, regulation of transcription, functions of non-coding RNA, organization of centromere and telomere regions of chromosomes, avian and amphibian oogenesis, germinal vesicles, lampbrush chromosomes, centromere-associated protein bodies, karyosphere formation, karyotype evolution.
Recent Honours
2008 - the Award for the best oral talk on the 18th European Colloquium on Animal Cytogenetics and Gene Mapping (Bucharest, Romany)
2009-2010 - Research Grant from the President of Russia for young scientists No.1.11.328.2009
2010 - the Award from the Saint-Petersburg Government for the scientific and educational achievements (Diploma No. 10027)
2011 - National L'Oreal-UNESCO fellowship within the programme "For Women in Science"
2014-2015 - Research Grant from the President of Russia for young scientists No.1.11.337.2014
2015 - Leonhard Euler Prize for outstanding scientific achievements in the field of Science and Technology from the Government of St. - Petersburg and St. - Petersburg Scientific Center RAS
2016 - Saint-Petersburg State University Award "Young scientists' contribution to science" for the scientific publications on the genome architecture and functional compartmentalization
of oocyte and somatic cell nucleus
Current research interests
Our current research is focusing on the investigation of functional compartmentalization and 3D-architecture of the oocyte nucleus (germinal vesicle). Nowadays, the role of intranuclear actin in the regulation of transcription and spatial genome organization in amphibian and avian growing oocytes has been studied in details. Molecular composition and 3-dimentional organization of meiotic chromosomes and intranuclear domains, such as amplified nucleoli and Cajal bodies, have been investigated by using of microinjections of fluorescently labeled molecules, modern techniques of confocal laser scanning microscopy and computational image-processing procedures, such as iso-surfacing and building the 3D-reconstructions. We also pay a special attention to establishment a link between the formation of various intranuclear domains and the expression of long non-coding RNA molecules. Giant lampbrush chromosomes from growing oocytes are been using as an adequate tool for exploration of genome expression. Fate and functions of non-coding RNA transcripts accumulating during the lampbrush stage of oogenesis are also under our investigation.
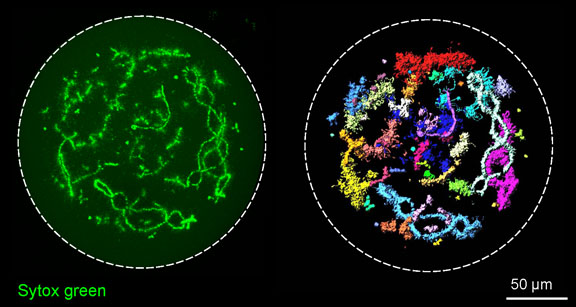
3D reconstruction of the giant nucleus isolated from growing oocyte of domestic chicken (Gallus gallus domesticus), scanned by using of confocal fluorescent microscope Leica TCS SP5 (Alla Krasikova, Antonina Maslova).
Other Qualifications
2005 - Practical Course on DNA-Microarrays Application in Comparative Cytogenetics, Leiden, Netherlands
2012 - "ENGAGE/GEUVADIS RNA-seq Workshop", European Bioinformatics Institute, Cambridge, UK
2013 - "Laboratory Management Course for Group Leaders", European Molecular Biology Organization, Heidelberg, Germany
2013 - "Fifth International Course on Fluorescence in situ Hybridization (FISH+array-CGH+microdissection)", University of Yena, Germany
2014 - "Writing in the Sciences", Stanford University Online Courses
2019-2020 - "Blackboard Learn Basics for teachers", Saint-Petersburg State University Online Courses
2020 - "Digital teacher. Practices and tools for organizing effective distance learning", Academy of National Economy and Public Administration, Online Course
Interships
2021 - Institute of Cytology and Genetics, RAS, Novosibirsk, Russia
2013, 2015 - University of Yena, Germany. Collaboration with Thomas Liehr, PhD. Full financial support from inter-University exchange programm
2005 - University of Wurzburg, Germany. Collaboration with Robert Hock, PhD. Full financial support from University of Wurzburg
Teaching activity
Author's lecture course "Genome architecture and nuclear compartmentalization"
Lectures within the course "Presentation of scientific data and the art of preparing an application for a grant"
Author's lecture course "Functions of non-coding RNA"
Students and trainees: Elena Vasilevskaya, Tatiana Khodyuchenko, Antonina Maslova, Dmitrij Dedukh, Darya Popova
Practical course "Structural and molecular analysis of biological systems"
Current Research Grants
2009-2011 Federal Grant-in-Aid Program "Human Capital for Science and Education in Innovative Russia" (Governmental Contract No. P1299);
2010-2012 Federal Grant-in-Aid Program "Human Capital for Science and Education in Innovative Russia" (Governmental Contract No. P1367);
2011-2013 Federal Grant-in-Aid Program "Human Capital for Science and Education in Innovative Russia" (Grant # 8122);
2011-2013 Research Grant from the Saint-Petersburg University No. 1.38.66.2011;
2013-2015 Research Grant from the Saint-Petersburg University for leading scientists;
2014-2015 Research Grant from the President of Russia for young scientists No.1.11.337.2014;
2014-2016 Research Grant from the Russian Science Foundation (# 14-14-00131);
2015-2016 Research Grant from the Russian Foundation for Basic Research (# 15-34-2102)
2019 Publishing Grant from the Russian Foundation for Basic Research (# 978-5-288-05984-1)
2019-2022 Research Grant from the Russian Science Foundation (# 19-74-20075)
Other activities
2011 - 2017 Member of the Scientific Board of the Biological Faculty (St Petersburg State University) (http://www.bio.spbu.ru/science/sciencecom.php);
2014 - 2022 Member of the Editorial Board of the "Biological Communications" Journal (https://biocomm.spbu.ru/);
2013 - 2017 Chair of the Council of Young Scientists of Saint-Petersburg State University (http://english.spbu.ru/science-4/sovetmu);
2015 - 2017 Member of the youth section of the Council for Science under the Ministry of Education and Science of the Russian Federation;
2015 - up to now Member of the European Cytogeneticists Association (http://www.e-c-a.eu/en/)
2017 - 2022 Deputy Editor-in-Chief of the "Biological Communications" Journal (https://biocomm.spbu.ru/);
2021 - up to now Associate Editor in Genome Organization and Dynamics (specialty section of Frontiers in Molecular Biosciences)
Major Publications


- A. Krasikova, T. Kulikova, M. Schelkunov, N. Makarova, A. Fedotova, V. Plotnikov, V. Berngardt, A. Maslova, A. Fedorov.
The first chicken oocyte nucleus whole transcriptomic profile defines the spectrum of maternal mRNA and non-coding RNA genes transcribed by the lampbrush chromosomes.Nucleic Acids Research (2024). DOI:10.1093/nar/gkae941
Full text
- Dedukh D., Maslova A., Al-Rikabi A., Padutsch N., Liehr T., Krasikova A.
Karyotypes of water frogs from the Pelophylax esculentus complex:
results of cross-species chromosomal painting. Chromosoma (2023). DOI:10.1007/s00412-023-00812-8
Full text
- Krasikova, A., Kulikova, T., Rodriguez Ramos, J.S., Maslova, A.
Assignment of the somatic A/B compartments to chromatin domains in giant transcriptionally active lampbrush chromosomes.
Epigenetics & Chromatin 16, 24 (2023). DOI: 10.1186/s13072-023-00499-2.
Full text
- Krasikova, A., Fishman V., Kulikova T.
Lampbrush chromosome studies in the post-genomic era.
Bioessays: News and Reviews in Molecular, Cellular and Developmental Biology. (2023) DOI:10.1002/bies.202200250
Full text
- Maslova A., Plotnikov V., Nuriddinov M. Gridina M., Fishman V.,Krasikova, A.
Hi-C analysis of genomic contacts revealed karyotype abnormalities in chicken HD3 cell line.
BMC Genomics. (2023) DOI:10.1186/s12864-023-09158-y
Full text
- Nurislamov, A.; Lagunov, T.; Gridina, M.; Krasikova, A.; Fishman, V.
Single-cell DNA methylation analysis of chicken lampbrush chromosomes.
International Journal of Molecular Sciences. (2022) DOI:10.3390/ijms232012601
Full text
- Gridina M., Taskina A., Lagunov T., Nurislamov A., Kulikova T., Krasikova A., Fishman V.
Comparison and critical assessment of single-cell Hi-C protocols.
Heliyon. (2022) DOI:10.1016/j.heliyon.2022.e11023
Full text
- Kulikova T., Maslova A., Starshova P., Rodriguez Ramos J.S., Krasikova A.
Comparison of the somatic TADs and lampbrush chromomere-loop complexes in transcriptionally active prophase I oocytes.
Chromosoma. (2022) DOI:10.1007/s00412-022-00780-5.
Full text
- Kulikova, T., Krasikova, A.
RNA-FISH on Lampbrush Chromosomes: Visualization of Individual Transcription Units
In: Cytogenetics and Molecular Cytogenetics, T. Liehr (Editor), CRC Press, Boca Raton, 2022, ISBN: 978-1003223658, (pp. 307-317).
Full text
- Dedukh D., Riumin S., Kolenda K., Chmielewska M., Rozenblut-Koscisty B., Kazmierczak M., Ogielska M., Krasikova A.
Maintenance of pure hybridogenetic water frog populations: Genotypic variability in progeny of diploid and triploid parents.
PLOS ONE 17(7):e0268574 (2022) DOI:10.1371/journal.pone.0268574.
Full text
- A. Maslova, Krasikova A.
FISH going meso-scale: a microscopic search for chromatin domains.
Frontiers in Cell and Developmental Biology.9:753097 (2021) DOI: 10.3389/fcell.2021.753097
Full text
- Dedukh D., Krasikova A.
Delete and survive: strategies of programmed genetic material elimination in eukaryotes.
Biological Reviews. 2021 DOI:10.1111/brv.12796
Full text
- Dedukh D, Riumin S, Chmielewska M, Rozenblut-Koscisty B, Kolenda K, Kazmierczak M, Dudzik A, Ogielska M, Krasikova A.
Micronuclei in germ cells of hybrid frogs from Pelophylax esculentus complex contain gradually eliminated chromosomes.
Scientific Reports. 10, 8720 (2020) DOI: 10.1038/s41598-020-64977-3
Full text
- Zlotina A., Maslova A., Pavlova O., Kosyakova N., Al-Rikabi A.,Liehr T., Krasikova A.
Insights Into Chromomere Organization Provided by Lampbrush Chromosome Microdissection and High-Throughput Sequencing.
Frontiers in Genetics. 11:57 (2020) DOI: 10.3389/fgene.2020.00057
Full text
- Kulikova T, Surkova A, Zlotina A, Krasikova A.
Mapping epigenetic modifications on chicken lampbrush chromosomes.
Full text
- Krasikova A.V. Kulikova T.V.
Lampbrush chromosomes: current concepts and research perspectives.
St Petersburg University Publishing House, 2019. 104 p. ISBN 978-5-288-05984-1 [In Russian].
Read more
- Dedukh D., Litvinchuk J., Svinin A., Litvinchuk S., Rosanov J.and Krasikova A.
Variation in hybridogenetic hybrid emergence between populations of water frogs from the Pelophylax esculentus complex.
PloS One. 14(11) (2019) DOI: 10.1371/journal.pone.0224759
Full text
- Krasikova A, Kulikova T.
Identification of Genomic Loci Responsible for the Formation of Nuclear Domains Using Lampbrush Chromosomes.
Noncoding RNA. 6(1):1 (2019) DOI: 10.3390/ncrna6010001
Full text
- Fishman, V., Battulin, N., Nuriddinov, M., Maslova, A., Zlotina, A., Strunov, A., Chervyakova, D., Korablev, A., Serov, O., Krasikova, A.
3D organization of chicken genome demonstrates evolutionary conservation of topologically associated domains and highlights unique architecture of erythrocytes' chromatin.
Nucleic Acids Research. DOI: 10.1093/nar/gky1103 (2018)
Full text
- Chmielewska, M., Dedukh, D., Haczkiewicz, K., Rozenblut-Kościsty, B., Kaźmierczak, M., Kolenda, K., Serwa, E., Pietras-Lebioda, A., Krasikova, A., Ogielska, M.
The programmed DNA elimination and formation of micronuclei in germ line cells of the natural hybridogenetic water frog Pelophylax esculentus.
Scientific Reports, 8 (1) 7870 (2018)
Full text
- Zlotina A., Dedukh D., Krasikova A.
Amphibian and Avian Karyotype Evolution: Insights from Lampbrush Chromosome Studies.
Genes (2017). V.8(11). doi: 10.3390/genes8110311
Full text
- D. Dedukh, S. Litvinchuk, J. Rosanov, D. Shabanov, A. Krasikova
Mutual maintenance of di- and triploid Pelophylax esculentus hybrids in R-E systems: results from artificial crossings experiments.
BMC Evolutionary Biology 17:220 (2017).
Full text
- A. V. Krasikova, T. V. Kulikova
Distribution of heterochromatin markers in lampbrush chromosomes in birds.
Russian Journal of Genetics 53(9): 1022–1029 (2017)
Full text
- Dedukh, D.V., Krasikova, A.V.
Methodological approaches for studying the european water frog Pelophylax esculentus complex.
Russ J Genet (2017) 53: 843.
- Kulikova T., Khodychenko T., Petrov Y., Krasikova A.
Low-voltage scanning electron microscopy study of lampbrush chromosomes and nuclear bodies in avian and amphibian oocytes.
Scientific Reports - 2016.- V.6 (36878). - doi: 10.1038/srep36878
Free full text
- I. Trofimova, A. Krasikova.
Transcription of highly repetitive tandemly organized DNA in amphibians and birds: a historical overview and modern concepts.
RNA Biology, 2016. DOI: 10.1080/15476286.2016.1240142
Free full text
- A.Zlotina, A. Krasikova.
FISH in Lampbrush Chromosomes.
In: Fluorescence in situ Hybridization (FISH)-Application Guide, T Liehr (Editor), 2nd Ed., Springer, Berlin, 2017, ISBN: 978-3662529577, pp. 445-458.
- Krasikova A., Kulikova T.
Atomic Force Microscopy of the avian and amphibian lampbrush chromosome.
The Russian Journal of Ornithology. - 2016. - V. 25 (1338) - P. 3457-3463.
Free full text [In Russian]
- Krasikova A., Kulikova T., Zlotina A.
Lampbrush chromosomes as a model for exploration into loci of nuclear domains formation.
Cell and Tissue Biology. - 2016. - V. 58 (4) - P. 277-280.
- Zlotina A., Kulikova T., Kosyakova N., Liehr T., Krasikova A.
Microdissection of lampbrush chromosomes as an approach for generation of locus-specific FISH-probes and samples for high-throughput sequencing.
BMC Genomics. 2016. 17:126
DOI: 10.1186/s12864-016-2437-4
Free full text
- Kulikova T., Chervyakova D., Zlotina A., Krasikova A., Gaginskaya E.
Giant poly (A)-rich RNP aggregates form at terminal regions of avian lampbrush chromosomes.
Chromosoma. 2015. 1-16
DOI:10.1007/s00412-015-0563-4
- Trofimova I., Chervyakova D., Krasikova A.
Transcription of subtelomere tandemly repetitive DNA in chicken embryogenesis.
Chromosome Research. 2015.
DOI: 10.1007/s10577-015-9487-3.
Free full text
- Maslova A., Zlotina A., Kosyakova N., Sidorova M., Krasikova A.
Three-dimensional architecture of tandem repeats in chicken interphase nucleus.
Chromosome Research. 2015.
DOI: 10.1007/s10577-015-9485-5.
Free full text
- Krasikova A., Fedorov A.
Expression Profiles of Long and Short RNAs in the Cytoplasm and Nuclei of Growing Chicken (Gallus gallus domesticus) Oocytes.
Russian Journal of Genetics. Applied Research - 2015. - V. 6(3) - P. 307-313.
Free full text
- Dedukh D., Litvinchuk S., Rosanov J., Mazepa G., Saifitdinova A., Shabanov D., Krasikova A.
Optional Endoreplication and Selective Elimination of Parental Genomes during Oogenesis in Diploid and Triploid Hybrid European Water Frogs.
PLoS ONE. 2015. V. 10(4).
Free full text
- Trofimova I., Popova D., Vasilevskaya E., Krasikova A.
Non-coding RNA derived from a conservative subtelomeric tandem repeat in chicken and Japanese quail somatic cells.
Molecular Cytogenetics. 2014. V. 7:102. P. 1-13
Free full text
- Khodyuchenko T., Krasikova A.
Cajal bodies and histone locus bodies: Molecular composition and function.
Russian Journal of Developmental Biology. 2014. V. 45:6. P. 297 312
Abstract
- Dedukh D., Mazepa G., Shabanov D., Rosanov J., Litvinchuk S., Borkin L., Saifitdinova A., Krasikova A.
Cytological maps of lampbrush chromosomes of European water frogs (Pelophylax esculentus complex) from the Eastern Ukraine.
BMC Genetics. 2013. V. 14. 26
Free full text
- Maslova A., Krasikova A.
Nuclear actin depolymerization in transcriptionally active avian and amphibian oocytes leads to collapse of intranuclear structures.
Nucleus. 2012. V. 3(3). P. 300-311
Abstract Free full text
- Zlotina A., Galkina S., Krasikova A., Crooijmans R.P., Groenen M.A., Gaginskaya E., Deryusheva S.
Centromere positions in chicken and Japanese quail chromosomes: de novo centromere formation versus pericentric inversions.
Chromosome Res. 2012. V. 20 (8). P. 1017-1032. DOI: 10.1007/s10577-012-9319-7
Abstract Free full text
- Krasikova A., Fukagawa T., Zlotina A.
High-resolution mapping and transcriptional activity analysis of chicken centromere sequences on giant lampbrush chromosomes.
Chromosome Res. 2012. V. 20 (8). P. 995-1008. DOI: 10.1007/s10577-012-9321-0
Abstract Free full text
- Krasikova A., Khodyuchenko T., Maslova A., Vasilevskaya E.
Three-dimensional organisation of RNA-processing machinery in avian growing oocyte nucleus.
Chromosome Res. 2012. V. 20 (8). P. 979-994. DOI: 10.1007/s10577-012-9321-0
Abstract Free full text
- Khodyuchenko T., Gaginskaya E., Krasikova A.
Non-canonical Cajal bodies form in the nucleus of late stage avian oocytes lacking functional nucleolus.
Histochem Cell Biol. 2012. V 138(1), P. 57-73
Abstract
- Maslova A., Krasikova A.
Spatial Arrangement of Macro-, Midi-, and Microchromosomes in Transcriptionally Active Nuclei of Growing Oocytes in Birds of Order Galliformes.
Cell and Tissue Biology. 2011. V. 5. N. 3. P. 281-293.
Abstract
- Krasikova A., Vasilevskaya E., Gaginskaya E.
Chicken Lampbrush Chromosomes: Transcription of Tandemly Repetitive DNA Sequences.
Russian Journal of Genetics. 2010. V. 46(10). P. 1173-1177.
Abstract
- Daks A., Deryusheva S., Krasikova A., Zlotina A., Gaginskaya E., Galkina S.
Lampbrush Chromosomes of the Japanese Quail (Coturnix coturnix japonica): A New Version of Cytogenetic Maps.
Russian Journal of Genetics. 2010. V. 46(10). P. 1178-1181.
Abstract
- Krasikova A., Gaginskaya E.
Organisation of centromere regions of chromosomes in the lampbrush phase.
Tsitologiia. 2010. V. 52(7). P. 515-33.
Abstract
- Zlotina A., Galkina S., Krasikova A., Crooijmans R.P., Groenen M.A., Gaginskaya E., Deryusheva S.
Precise centromere positioning on chicken chromosome 3.
Cytogenet Genome Res. 2010. V. 129(4). P.310-313.
Abstract
- Krasikova A., Daks A., Zlotina A., Gaginskaya E.
Polymorphic heterochromatic segments in Japanese quail microchromosomes.
Cytogenetics and Genome Research. 2009. V. 126. P.148-155
Abstract
- Gaginskaya E., Kulikova T., Krasikova A.
Avian Lampbrush Chromosomes: a Powerful Tool for Exploration of Genome Expression.
Cytogenet Genome Res. 2009. V.124. P.251-267.
Abstract
- Deryusheva S., Krasikova A., Kulikova T., Gaginskaya E.
Tandem 41-bp repeats in chicken and Japanese quail genomes: FISH mapping and transcription analysis on lampbrush chromosomes.
Chromosoma. 2007. V. 116 P.519-530.
Abstract
- Krasikova A., Deryusheva S., Galkina S., Kurganova A., Evteev A., Gaginskaya E.
On the positions of centromeres in chicken lampbrush chromosomes.
Chromosome Res. 2006 V. 14 P.777-789.
Abstract
- Krasikova A., Barbero J.L., Gaginskaya E.
Cohesion proteins are present in centromere protein bodies associated with avian lampbrush chromosomes.
Chromosome Res. 2005. V.13. P.675-685.
Abstract
- Krasikova A., Kulikova T., Saifitdinova A., Derjusheva S., Gaginskaya E.
Centromeric protein bodies on avian lampbrush chromosomes contain a protein detectable with an antibody against DNA topoisomerase II.
Chromosoma. 2004. V. 116. P. 316 323.
Abstract
- Derjusheva S., Kurganova A., Krasikova A., Saifitdinova A., Habermann F.A., Gaginskaya E.
Precize identification of chicken chromosomes in the lampbrush form using chromosome painting probes.
Chromosome research. 2003. V. 11. P. 749 757.
Abstract
- Saifitdinova A., Derjusheva S., Krasikova A., Gaginskaya E.
Lampbrush chromosomes of the chaffinch ( Fringilla coelebs L.).
Chromosome research. 2003. V. 11. P. 99-113.
Abstract




